Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with the Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.1 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the Microorganisms.
- Companion journal: Applied Microbiology.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Transcriptome Analysis of mfs2-Defective Penicillium digitatum Mutant to Reveal Importance of Pdmfs2 in Developing Fungal Prochloraz Resistance
Microorganisms 2024, 12(5), 888; https://doi.org/10.3390/microorganisms12050888 (registering DOI) - 28 Apr 2024
Abstract
Demethylation inhibitors (DMIs), including prochloraz, are popular fungicides to control citrus postharvest pathogens such as Penicillium digitatum (green mold). However, many P. digitatum strains have developed prochloraz resistance, which decreases drug efficacy. Specific major facilitator superfamily (MFS) transporter gene mfs2, encoding drug-efflux
[...] Read more.
Demethylation inhibitors (DMIs), including prochloraz, are popular fungicides to control citrus postharvest pathogens such as Penicillium digitatum (green mold). However, many P. digitatum strains have developed prochloraz resistance, which decreases drug efficacy. Specific major facilitator superfamily (MFS) transporter gene mfs2, encoding drug-efflux pump protein MFS2, has been identified in P. digitatum strain F6 (PdF6) to confer fungal strain prochloraz resistance. However, except for the drug-efflux pump function of MFS2, other mechanisms relating to the Pdmfs2 are not fully clear. The present study reported a transcriptome investigation on the mfs2-defective P. digitatum strain. Comparing to the wild-type strain, the mfs2-defective strain showed 717 differentially expressed genes (DEGs) without prochloraz induction, and 1221 DEGs with prochloraz induction. The obtained DEGs included multiple isoforms of MFS transporter-encoding genes, ATP-binding cassette (ABC) transporter-encoding genes, and multidrug and toxic compound extrusion (MATE) family protein-encoding genes. Many of these putative drug-efflux pump protein-encoding genes had significantly lower transcript abundances in the mfs2-defective P. digitatum strain at prochloraz induction, as compared to the wild-type strain, including twenty-two MFS transporter-encoding genes (MFS1 to MFS22), two ABC transporter-encoding genes (ABC1 and ABC2), and three MATE protein-encoding genes (MATE1 to MATE3). The prochloraz induction on special drug-efflux pump protein genes in the wild-type strain was not observed in the mfs2-defective strain, including MFS21, MFS22, ABC2, MATE1, MATE2, and MATE3. On the other hand, the up-regulation of other drug-efflux pump protein genes in the mfs2-defective strain cannot recover the fungal prochloraz resistance, including MFS23, MFS26, MFS27, MFS31, MFS33, and ABC3 to ABC8. The functional enrichment of DEGs based on Kyoto Encyclopedia of Genes and Genomes (KEGG), Clusters of Orthologous Groups (COG), and euKaryotic Orthologous Groups (KOG) database resources suggested some essential contributors to the mfs2-relating prochloraz resistance, including ribosome biosynthesis-related genes, oxidative phosphorylation genes, steroid biosynthesis-related genes, fatty acid and lipid metabolism-related genes, and carbon- and nitrogen-metabolism-related genes. The results indicated that the MFS2 transporter might be involved in the regulation of multiple drug-efflux pump protein gene expressions and multiple metabolism-related gene expressions, thus playing an important role in developing P. digitatum prochloraz resistance.
Full article
(This article belongs to the Special Issue Fungicide Resistance in Plant Pathogen)
Open AccessArticle
Managing Super Pests: Interplay between Pathogens and Symbionts Informs Biocontrol of Whiteflies
by
Weili Yan, Saixian Wang, Jialei Liu, Dan Zhai, Hang Lu, Jingjing Li, Rune Bai, Caiyan Lei, Luyang Song, Chenchen Zhao and Fengming Yan
Microorganisms 2024, 12(5), 887; https://doi.org/10.3390/microorganisms12050887 (registering DOI) - 28 Apr 2024
Abstract
Bemisia tabaci is distributed globally and incurs considerable economic and ecological costs as an agricultural pest and viral vector. The entomopathogenic fungus Metarhizium anisopliae has been known for its insecticidal activity, but its impacts on whiteflies are understudied. We investigated how infection with
[...] Read more.
Bemisia tabaci is distributed globally and incurs considerable economic and ecological costs as an agricultural pest and viral vector. The entomopathogenic fungus Metarhizium anisopliae has been known for its insecticidal activity, but its impacts on whiteflies are understudied. We investigated how infection with the semi-persistently transmitted Cucurbit chlorotic yellows virus (CCYV) affects whitefly susceptibility to M. anisopliae exposure. We discovered that viruliferous whiteflies exhibited increased mortality when fungus infection was present compared to non-viruliferous insects. High throughput 16S rRNA sequencing also revealed significant alterations of the whitefly bacterial microbiome diversity and structure due to both CCYV and fungal presence. Specifically, the obligate symbiont Portiera decreased in relative abundance in viruliferous whiteflies exposed to M. anisopliae. Facultative Hamiltonella and Rickettsia symbionts exhibited variability across groups but dominated in fungus-treated non-viruliferous whiteflies. Our results illuminate triangular interplay between pest insects, their pathogens, and symbionts—dynamics which can inform integrated management strategies leveraging biopesticides This work underscores the promise of M. anisopliae for sustainable whitefly control while laying the groundwork for elucidating mechanisms behind microbe-mediated shifts in vector competence.
Full article
(This article belongs to the Special Issue Plant Pathogens: Monitoring, Identification and Biological Control)
Open AccessReview
What do We Know about Cryptic Aspergillosis?
by
Nicholas Geremia, Federico Giovagnorio, Agnese Colpani, Andrea De Vito, Giorgia Caruana, Maria Chiara Meloni, Giordano Madeddu, Sandro Panese and Saverio Giuseppe Parisi
Microorganisms 2024, 12(5), 886; https://doi.org/10.3390/microorganisms12050886 (registering DOI) - 28 Apr 2024
Abstract
Cryptic Aspergillus species are increasingly recognized as pathogens involved in human disease. They are ubiquitarian fungi with high tenacity in their environment and can express various resistance mechanisms, often due to exposure to antifungal agents employed in agriculture and farming. The identification of
[...] Read more.
Cryptic Aspergillus species are increasingly recognized as pathogens involved in human disease. They are ubiquitarian fungi with high tenacity in their environment and can express various resistance mechanisms, often due to exposure to antifungal agents employed in agriculture and farming. The identification of such species is increasing thanks to molecular techniques, and a better description of this type of pathogen is granted. Nevertheless, the number of species and their importance in the clinical setting still need to be well studied. Furthermore, their cross-sectional involvement in animal disease, plants, and human activities requires a multidisciplinary approach involving experts from various fields. This comprehensive review aims to provide a sharp vision of the cryptic Aspergillus species, from the importance of correct identification to the better management of the infections caused by these pathogens. The review also accentuates the importance of the One Health approach for this kind of microorganism, given the interconnection between environmental exposure and aspergillosis, embracing transversely the multidisciplinary process for managing the cryptic Aspergillus species. The paper advocates the need for improving knowledge in this little-known species, given the burden of economic and health implications related to the diffusion of these bugs.
Full article
(This article belongs to the Special Issue Emerging Pathogens in the Context of One Health)
►▼
Show Figures
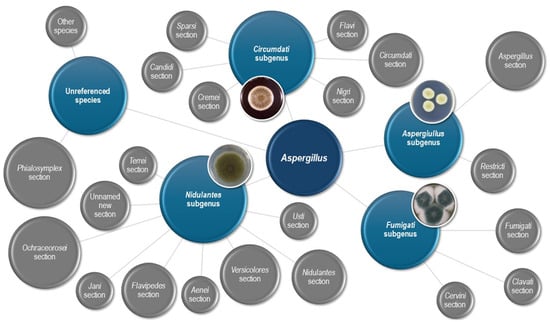
Figure 1
Open AccessReview
Paulownia Witches’ Broom Disease: A Comprehensive Review
by
Yajie Zhang, Zesen Qiao, Jidong Li and Assunta Bertaccini
Microorganisms 2024, 12(5), 885; https://doi.org/10.3390/microorganisms12050885 (registering DOI) - 28 Apr 2024
Abstract
Phytoplasmas are insect-transmitted bacterial pathogens associated with diseases in a wide range of host plants, resulting in significant economic and ecological losses. Perennial deciduous trees in the genus Paulownia are widely planted for wood harvesting and ornamental purposes. Paulownia witches’ broom (PaWB) disease,
[...] Read more.
Phytoplasmas are insect-transmitted bacterial pathogens associated with diseases in a wide range of host plants, resulting in significant economic and ecological losses. Perennial deciduous trees in the genus Paulownia are widely planted for wood harvesting and ornamental purposes. Paulownia witches’ broom (PaWB) disease, associated with a 16SrI-D subgroup phytoplasma, is a destructive disease of paulownia in East Asia. The PaWB phytoplasmas are mainly transmitted by insect vectors in the Pentatomidae (stink bugs), Miridae (mirid bugs) and Cicadellidae (leafhoppers) families. Diseased trees show typical symptoms, such as branch and shoot proliferation, which together are referred to as witches’ broom. The phytoplasma presence affects the physiological and anatomical structures of paulownia. Gene expression in paulownia responding to phytoplasma presence have been studied at the transcriptional, post-transcriptional, translational and post-translational levels by high throughput sequencing techniques. A PaWB pathogenic mechanism frame diagram on molecular level is summarized. Studies on the interactions among the phytoplasma, the insect vectors and the plant host, including the mechanisms underlying how paulownia effectors modify processes of gene expression, will lead to a deeper understanding of the pathogenic mechanisms and to the development of efficient control measures.
Full article
(This article belongs to the Special Issue Phloem Localized Insect Transmitted Bacteria Associated with Plant Diseases)
Open AccessReview
Addressing Sexually Transmitted Infections Due to Neisseria gonorrhoeae in the Present and Future
by
Julia Colón Pérez, Rosa-Antía Villarino Fernández, Adrián Domínguez Lago, María Mercedes Treviño Castellano, María Luisa Pérez del Molino Bernal, Sandra Sánchez Poza and Eva Torres-Sangiao
Microorganisms 2024, 12(5), 884; https://doi.org/10.3390/microorganisms12050884 (registering DOI) - 28 Apr 2024
Abstract
It was in the 1800s when the first public publications about the infection and treatment of gonorrhoea were released. However, the first prevention programmes were only published a hundred years later. In the 1940s, the concept of vaccination was introduced into clinical prevention
[...] Read more.
It was in the 1800s when the first public publications about the infection and treatment of gonorrhoea were released. However, the first prevention programmes were only published a hundred years later. In the 1940s, the concept of vaccination was introduced into clinical prevention programmes to address early sulphonamide resistance. Since then, tons of publications on Neisseria gonorrhoeae are undisputed, around 30,000 publications today. Currently, the situation seems to be just as it was in the last century, nothing has changed or improved. So, what are we doing wrong? And more importantly, what might we do? The review presented here aims to review the current situation regarding the resistance mechanisms, prevention programmes, treatments, and vaccines, with the challenge of better understanding this special pathogen. The authors have reviewed the last five years of advancements, knowledge, and perspectives for addressing the Neisseria gonorrhoeae issue, focusing on new therapeutic alternatives.
Full article
(This article belongs to the Special Issue Infectious Diseases: New Approaches to Old Problems 3.0)
►▼
Show Figures
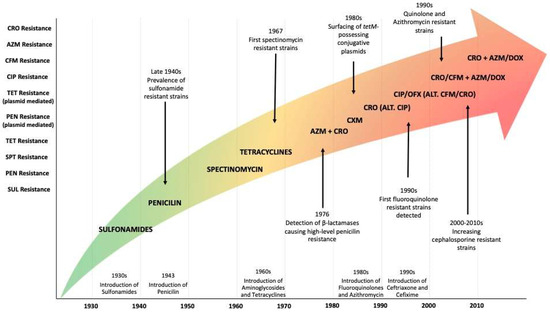
Figure 1
Open AccessArticle
Genetic Analyses of Rare ESBL ST628 Klebsiella pneumoniae Detected during a Protracted Nosocomial Outbreak in the United Kingdom
by
Stephen Mark Edward Fordham, Francis Drobniewski, Magdalena Barrow, Melissa Hutchings, Kate Crowther, Denise Richards, Paul Bolton, Anna Mantzouratou and Elizabeth Sheridan
Microorganisms 2024, 12(5), 883; https://doi.org/10.3390/microorganisms12050883 (registering DOI) - 28 Apr 2024
Abstract
Klebsiella pneumoniae (K. pneumoniae) cultures from a hospital-wide outbreak in the UK, which lasted for over 12 months, were sequenced. We sought to sequence and genetically characterise the outbreak strain. Antibiotic Susceptibility Testing (AST) was performed on 65 K. pneumoniae isolates
[...] Read more.
Klebsiella pneumoniae (K. pneumoniae) cultures from a hospital-wide outbreak in the UK, which lasted for over 12 months, were sequenced. We sought to sequence and genetically characterise the outbreak strain. Antibiotic Susceptibility Testing (AST) was performed on 65 K. pneumoniae isolates saved from the outbreak. All isolates were sequenced using the Oxford Nanopore Technologies (ONT) MinION flowcell: 10 isolates, including the isolate with the earliest collection date in 2017, were additionally sequenced on the NovaSeq 6000 platform to build high-accuracy nanopore-illumina assemblies. Among the sequenced strains, 60 were typed as ST628. 96.6% (n = 58/60) ST628 strains harboured a large ~247-kb FIB(K) plasmid carrying up to 11 antimicrobial resistance genes, including the extended-spectrum beta-lactamase (ESBL) gene, blaCTX-M-15. Clonality between the outbreak isolates was confirmed using single nucleotide polymorphism (SNP) typing. The outbreak strains were phylogenetically related to clinical ST628 strains identified in 2012, 6 years prior to the outbreak. A rare ESBL K. pneumoniae K2 ST628 strain harbouring a multi-drug resistant (MDR) plasmid encoding the ESBL gene blaCTX-M-15 was detected across multiple independent wards during the protracted nosocomial outbreak. Surveillance of this strain is recommended to prevent future nosocomial outbreaks.
Full article
(This article belongs to the Section Systems Microbiology)
►▼
Show Figures
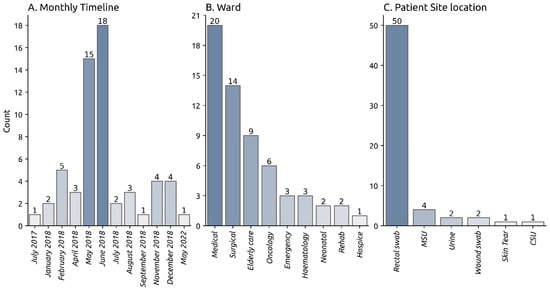
Figure 1
Open AccessArticle
Advancements in Fermented Beverage Safety: Isolation and Application of Clavispora lusitaniae Cl-p for Ethyl Carbamate Degradation and Enhanced Flavor Profile
by
Yingchun Zhao, Jun Liu, Han Wang, Fayuan Gou, Yiwei He and Lijuan Yang
Microorganisms 2024, 12(5), 882; https://doi.org/10.3390/microorganisms12050882 (registering DOI) - 28 Apr 2024
Abstract
Ethyl carbamate (EC) is a natural by-product in the production of fermented food and alcoholic beverages and is carcinogenic and genotoxic, posing a significant food safety concern. In this study, Clavispora lusitaniae Cl-p with a strong EC degradation ability was isolated from Daqu
[...] Read more.
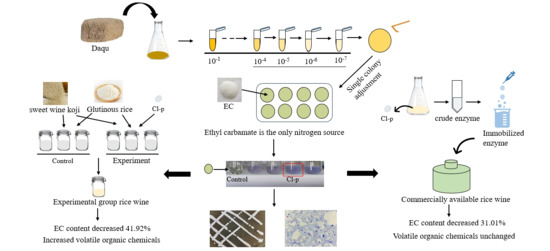
Ethyl carbamate (EC) is a natural by-product in the production of fermented food and alcoholic beverages and is carcinogenic and genotoxic, posing a significant food safety concern. In this study, Clavispora lusitaniae Cl-p with a strong EC degradation ability was isolated from Daqu rich in microorganisms by using EC as the sole nitrogen source. When 2.5 g/L of EC was added to the fermentation medium, the strain decomposed 47.69% of ethyl carbamate after five days of fermentation. It was unexpectedly found that the strain had the ability to produce aroma and ester, and the esterification power reached 30.78 mg/(g·100 h). When the strain was added to rice wine fermentation, compared with the control group, the EC content decreased by 41.82%, and flavor substances such as ethyl acetate and β-phenylethanol were added. The EC degradation rate of the immobilized crude enzyme in the finished yellow rice wine reached 31.01%, and the flavor substances of yellow rice wine were not affected. The strain is expected to be used in the fermented food industry to reduce EC residue and improve the safety of fermented food.
Full article
(This article belongs to the Section Food Microbiology)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Optimizing Akkermansia muciniphila Isolation and Cultivation: Insights into Gut Microbiota Composition and Potential Growth Promoters in a Chinese Cohort
by
Xiangyu Meng, Chen Xv, Jiaping Lv, Shuwen Zhang, Changlu Ma and Xiaoyang Pang
Microorganisms 2024, 12(5), 881; https://doi.org/10.3390/microorganisms12050881 (registering DOI) - 28 Apr 2024
Abstract
The study aims to analyze the composition of the gut microbiota in Chinese individuals using metagenomic sequencing technology, with a particular focus on the abundance of Akkermansia muciniphila (Akk). To improve the efficiency of Akk isolation and identification accuracy, modifications were made to
[...] Read more.
The study aims to analyze the composition of the gut microbiota in Chinese individuals using metagenomic sequencing technology, with a particular focus on the abundance of Akkermansia muciniphila (Akk). To improve the efficiency of Akk isolation and identification accuracy, modifications were made to the enrichment culture medium and 16S rRNA universal primers. Additionally, potential growth-promoting factors that stimulate Akk growth were explored through in vitro screening. The research results revealed that the abundance of Akk in Chinese fecal samples ranged from 0.004% to 0.4%. During optimization, a type of animal protein peptide significantly enhanced the enrichment efficiency of Akk, resulting in the isolation of three Akk strains from 14 fecal samples. Furthermore, 17 different growth-promoting factors were compared, and four factors, including galactose, sialic acid, lactose, and chitosan, were identified as significantly promoting Akk growth. Through orthogonal experiments, the optimal ratio of these four growth-promoting factors was determined to be 1:1:2:1. After adding 1.25% of this growth-promoting factor combination to the standard culture medium, Akk was cultivated at 37° for 36 h, achieving an OD600nm value of 1.169, thus realizing efficient proliferation and optimized cultivation of Akk. This study provides important clues for a deeper understanding of the gut microbiota composition in Chinese individuals, while also offering effective methods for the isolation and cultivation of Akk, laying the groundwork for its functional and application research in the human body.
Full article
(This article belongs to the Special Issue Effects of Gut Microbiota on Human Health and Disease)
►▼
Show Figures
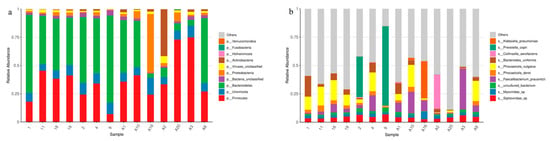
Figure 1
Open AccessArticle
Wine Barrel Biofilm as a Source of Yeasts with Non-Conventional Properties
by
Giorgia Perpetuini, Alessio Pio Rossetti, Arianna Rapagnetta, Giuseppe Arfelli, Roberta Prete and Rosanna Tofalo
Microorganisms 2024, 12(5), 880; https://doi.org/10.3390/microorganisms12050880 (registering DOI) - 27 Apr 2024
Abstract
This study investigated the main microbial groups characterizing the interior surface of oak barrels from different years (1890, 1895, 1920, 1975, 2008) used in the production of vino cotto. The yeasts were characterized for the following properties: γ-aminobutyric acid (GABA) production, antioxidant activity,
[...] Read more.
This study investigated the main microbial groups characterizing the interior surface of oak barrels from different years (1890, 1895, 1920, 1975, 2008) used in the production of vino cotto. The yeasts were characterized for the following properties: γ-aminobutyric acid (GABA) production, antioxidant activity, air–liquid interfacial biofilm formation, and anthocyanin adsorption capacity. Community-level physiological profile analysis revealed that the microbial communities inside the barrels used the tested carbon sources in different manners. The following yeast species were identified: Millerozyma farinosa, Zygosaccharomyces bisporus, Wickerhamiella versatilis, Zygosaccharomyces bailii, Starmerella lactis-condensi, and Zygosaccharomyces rouxii. All the strains were able to produce GABA, and S. lactis-condensi, Z. bisporus and Z. rouxii were the highest producers (more than 600 mg/L). The Z. rouxii and Z. bailii strains showed the highest antioxidant activity. Only seven strains out of ten M. farinosa formed air–liquid interfacial biofilm. None of the M. farinosa strains adsorbed anthocyanins on their cell wall. The other strains adsorbed anthocyanins in a strain-dependent way, and the highest adsorption was observed for the W. versatilis strains. The yeasts isolated in this study could be used to increase the functional properties and the quality of fermented foods and beverages.
Full article
(This article belongs to the Section Biofilm)
Open AccessReview
Transmission and Persistence of Infant Gut-Associated Bifidobacteria
by
Margaret A. Hilliard and David A. Sela
Microorganisms 2024, 12(5), 879; https://doi.org/10.3390/microorganisms12050879 (registering DOI) - 27 Apr 2024
Abstract
Bifidobacterium infantis are the primary colonizers of the infant gut, yet scientific research addressing the transmission of the genus Bifidobacterium to infants remains incomplete. This review examines microbial reservoirs of infant-type Bifidobacterium that potentially contribute to infant gut colonization. Accordingly, strain inheritance from
[...] Read more.
Bifidobacterium infantis are the primary colonizers of the infant gut, yet scientific research addressing the transmission of the genus Bifidobacterium to infants remains incomplete. This review examines microbial reservoirs of infant-type Bifidobacterium that potentially contribute to infant gut colonization. Accordingly, strain inheritance from mother to infant via the fecal-oral route is likely contingent on the bifidobacterial strain and phenotype, whereas transmission via the vaginal microbiota may be restricted to Bifidobacterium breve. Additional reservoirs include breastmilk, horizontal transfer from the environment, and potentially in utero transfer. Given that diet is a strong predictor of Bifidobacterium colonization in early life and the absence of Bifidobacterium is observed regardless of breastfeeding, it is likely that additional factors are responsible for bifidobacterial colonization early in life.
Full article
(This article belongs to the Special Issue Beneficial Microbes and Gastrointestinal Microbiota 2.0)
►▼
Show Figures
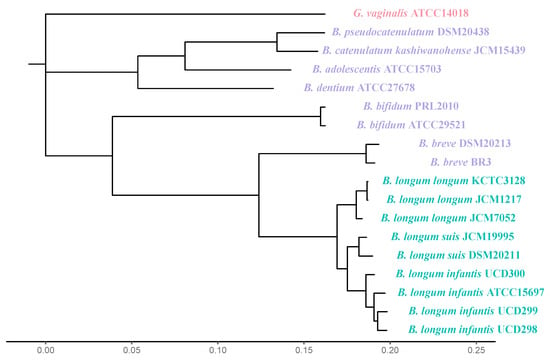
Figure 1
Open AccessArticle
New Species-Specific Real-Time PCR Assays for Colletotrichum Species Causing Bitter Rot of Apple
by
Diana J. McHenry and Srđan G. Aćimović
Microorganisms 2024, 12(5), 878; https://doi.org/10.3390/microorganisms12050878 (registering DOI) - 27 Apr 2024
Abstract
Bitter rot of apple is an economically important worldwide disease caused by different Colletotrichum species, depending on many factors such as climate, geography, other hosts, and crop management practices. Culture, morphology, and single-locus sequencing-based methods for identifying the Colletotrichum species are severely limited
[...] Read more.
Bitter rot of apple is an economically important worldwide disease caused by different Colletotrichum species, depending on many factors such as climate, geography, other hosts, and crop management practices. Culture, morphology, and single-locus sequencing-based methods for identifying the Colletotrichum species are severely limited in effectiveness, while the multilocus sequence typing methods available for delineating species are costly, time-intensive, and require high expertise. We developed species-specific hydrolysis probe real-time PCR assays for the following nine Colletotrichum species causing bitter rot in the Mid-Atlantic U.S.A.: C. fructicola, C. chrysophilum, C. noveboracense, C. gloeosporioides s.s., C. henanense, C. siamense and C. theobromicola from the C. gloeosporioides species complex, and C. fioriniae and C. nymphaeae from the C. acutatum species complex. After searching 14 gene regions, we designed primers and probes in 5 of them for the nine target species. Four primer–probe set pairs were able to be duplexed. Sensitivity tests showed as little as 0.5 pg DNA were detectable. These real-time PCR assays will provide rapid and reliable identification of these key Colletotrichum species and will be critically important for studies aiming to elucidate their biology, epidemiology, and management on apples as the number one produced and consumed tree fruit in the U.S.A.
Full article
(This article belongs to the Special Issue Colletotrichum Pathogens in Plants)
Open AccessArticle
Isolation of Vibrio cholerae and Vibrio vulnificus from Estuarine Waters, and Genotyping of V. vulnificus Isolates Using Loop-Mediated Isothermal Amplification
by
Shin-ichi Miyoshi, Megumi Kurata, Riho Hirose, Masaya Yoshikawa, Yong Liang, Yosuke Yamagishi and Tamaki Mizuno
Microorganisms 2024, 12(5), 877; https://doi.org/10.3390/microorganisms12050877 (registering DOI) - 27 Apr 2024
Abstract
Bacteria in the genus Vibrio are ubiquitous in estuarine and coastal waters. Some species (including Vibrio cholerae and Vibrio vulnificus) are known human pathogens causing ailments like cholera, diarrhea, or septicemia. Notably, V. vulnificus can also cause a severe systemic infection (known as
[...] Read more.
Bacteria in the genus Vibrio are ubiquitous in estuarine and coastal waters. Some species (including Vibrio cholerae and Vibrio vulnificus) are known human pathogens causing ailments like cholera, diarrhea, or septicemia. Notably, V. vulnificus can also cause a severe systemic infection (known as vibriosis) in eels raised in aquaculture facilities. Water samples were periodically collected from the estuary of the Asahi River, located in the southern part of Okayama City, Japan. These samples were directly plated onto CHROMagar Vibrio plates, and colonies displaying turquoise-blue coloration were selected. Thereafter, polymerase chain reaction was used to identify V. cholerae and V. vulnificus. A total of 30 V. cholerae strains and 194 V. vulnificus strains were isolated during the warm season when the water temperature (WT) was higher than 20 °C. Concurrently, an increase in coliforms was observed during this period. Notably, V. vulnificus has two genotypes, designated as genotype 1 and genotype 2. Genotype 1 is pathogenic to humans, while genotype 2 is pathogenic to both humans and eels. The loop-mediated isothermal amplification method was developed to rapidly determine genotypes at a low cost. Of the 194 strains isolated, 80 (41.2%) were identified as genotype 1 strains. Among the 41 strains isolated when the WTs were higher than 28 °C, 25 strains (61.0%) belonged to genotype 1. In contrast, of the 32 strains isolated when the WTs were lower than 24 °C, 27 strains (84.4%) belonged to genotype 2. These results suggest that the distribution of the two genotypes was influenced by WT.
Full article
(This article belongs to the Special Issue Water Microorganisms Associated with Human Health)
►▼
Show Figures
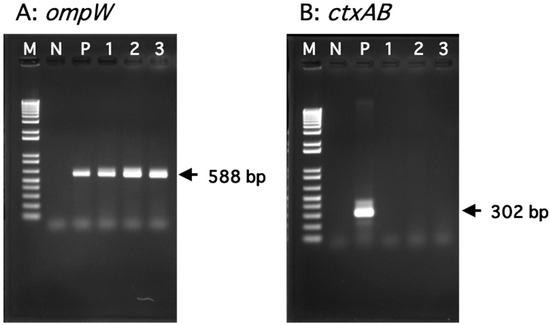
Figure 1
Open AccessArticle
Characterization of Escherichia coli Strains for Novel Production of Plasmodium ovale Lactate Dehydrogenase
by
Jae-Won Choi, Sang-Oh Ha, Yeon-Jun Kim, Jun-Seop Shin, Min-Ji Choi, Si-Eun Yu, Junghun Han, Eun-Ji Park, Kyoung Sik Park and Jung Hoon Kang
Microorganisms 2024, 12(5), 876; https://doi.org/10.3390/microorganisms12050876 (registering DOI) - 27 Apr 2024
Abstract
Malaria is one of the most prevalent diseases worldwide with high incidence and mortality. Among the five species that can infect humans, Plasmodium ovale morphologically resembles P. vivax, resulting in misidentification and confusion in diagnosis, and is responsible for malarial disease relapse
[...] Read more.
Malaria is one of the most prevalent diseases worldwide with high incidence and mortality. Among the five species that can infect humans, Plasmodium ovale morphologically resembles P. vivax, resulting in misidentification and confusion in diagnosis, and is responsible for malarial disease relapse due to the formation of hypnozoites. P. ovale receives relatively less attention compared to other major parasites, such as P. falciparum and P. vivax, primarily due to its lower pathogenicity, mortality rates, and prevalence rates. To efficiently produce lactate dehydrogenase (LDH), a major target for diagnosing malaria, this study used three Escherichia coli strains, BL21(DE3), BL21(DE3)pLysS, and Rosetta(DE3), commonly used for recombinant protein production. These strains were characterized to select the optimal strain for P. ovale LDH (PoLDH) production. Gene cloning for recombinant PoLDH production and transformation of the three strains for protein expression were performed. The optimal PoLDH overexpression and washing buffer conditions in nickel-based affinity chromatography were established to ensure high-purity PoLDH. The yields of PoLDH expressed by the three strains were as follows: BL21(DE3), 7.6 mg/L; BL21(DE3)pLysS, 7.4 mg/L; and Rosetta(DE3), 9.5 mg/L. These findings are expected to be highly useful for PoLDH-specific diagnosis and development of antimalarial therapeutics.
Full article
(This article belongs to the Special Issue Advances in Microbial Cell Factories, 2nd Edition)
Open AccessArticle
Ovine and Caprine Strains of Corynebacterium pseudotuberculosis on Czech Farms—A Comparative Study
by
Jirina Markova, Denisa Langova, Vladimir Babak and Iveta Kostovova
Microorganisms 2024, 12(5), 875; https://doi.org/10.3390/microorganisms12050875 (registering DOI) - 27 Apr 2024
Abstract
Caseous lymphadenitis (CLA) is a worldwide disease of small ruminants caused by Corynebacterium pseudotuberculosis, a facultative intracellular pathogen that is able to survive and multiply in certain white blood cells of the host. In this study, 33 strains of C. pseudotuberculosis were
[...] Read more.
Caseous lymphadenitis (CLA) is a worldwide disease of small ruminants caused by Corynebacterium pseudotuberculosis, a facultative intracellular pathogen that is able to survive and multiply in certain white blood cells of the host. In this study, 33 strains of C. pseudotuberculosis were isolated from sheep and goats suffering from CLA on nine farms in the Czech Republic. All these strains were tested for their antibiotic susceptibility, ability to form a biofilm and resistance to the effects of commonly used disinfectant agents. To better understand the virulence of C. pseudotuberculosis, the genomes of strains were sequenced and comparative genomic analysis was performed with another 123 genomes of the same species, including ovis and equi biovars, downloaded from the NCBI. The genetic determinants for the virulence factors responsible for adherence and virulence factors specialized for iron uptake and exotoxin phospholipase D were revealed in every analyzed genome. Carbohydrate-Active Enzymes were compared, revealing the presence of genetic determinants encoding exo-α-sialidase (GH33) and the CP40 protein in most of the analyzed genomes. Thirty-three Czech strains of C. pseudotuberculosis were identified as the biovar ovis on the basis of comparative genome analysis. All the compared genomes of the biovar ovis strains were highly similar regardless of their country of origin or host, reflecting their clonal behavior.
Full article
(This article belongs to the Special Issue Bacterial Infections and Antimicrobial Resistance in Animals)
►▼
Show Figures
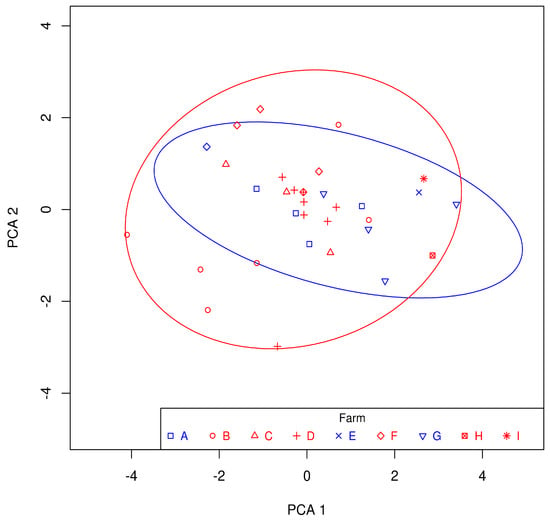
Figure 1
Open AccessCommunication
Microbial Community and Metabolome Analysis of the Porcine Intestinal Damage Model Induced by the IPEC-J2 Cell Culture-Adapted Porcine Deltacoronavirus (PDCoV) Infection
by
Ying Shi, Benqiang Li, Jinghua Cheng, Jie Tao, Pan Tang, Jiajie Jiao and Huili Liu
Microorganisms 2024, 12(5), 874; https://doi.org/10.3390/microorganisms12050874 (registering DOI) - 27 Apr 2024
Abstract
This study was conducted to elucidate the intestinal damage induced by the IPEC-J2 cell culture-passaged PDCoV. The results showed that PDCoV disrupted the intestinal structure and increased intestinal permeability, causing abnormalities in mucosal pathology. Additionally, PDCoV induced an imbalance in the intestinal flora
[...] Read more.
This study was conducted to elucidate the intestinal damage induced by the IPEC-J2 cell culture-passaged PDCoV. The results showed that PDCoV disrupted the intestinal structure and increased intestinal permeability, causing abnormalities in mucosal pathology. Additionally, PDCoV induced an imbalance in the intestinal flora and disturbed its stability. Microbial community profiling revealed bacterial enrichment (e.g., Proteobacteria) and reduction (e.g., Firmicutes and Bacteroidetes) in the PDCoV-inoculated piglet model. In addition, metabolomics analysis indicated that 82 named differential metabolites were successfully quantified, including 37 up-regulated and 45 down-regulated metabolites. Chenodeoxycholic acid, sphingosine, and oleanolic aldehyde levels were reduced in PDCoV-inoculated piglets, while phenylacetylglycine and geranylgeranyl-PP levels were elevated. Correlation analysis indicated a negative correlation between Escherichia-Shigella and choline, succinic acid, creatine, phenyllactate, and hippuric acid. Meanwhile, Escherichia-Shigella was positively correlated with acetylcholine, L-Glutamicacid, and N-Acetylmuramate. Roseburia, Lachnospiraceae_UCG-010, Blautia, and Limosilactobacillus were negatively and positively correlated with sphingosine, respectively. These data suggested PDCoV-inoculated piglets exhibited significant taxonomic perturbations in the gut microbiome, which may result in a significantly altered metabolomic profile.
Full article
(This article belongs to the Section Veterinary Microbiology)
Open AccessArticle
Lactiplantibacillus plantarum ST-III and Lacticaseibacillus rhamnosus KF7 Enhance the Intestinal Epithelial Barrier in a Dual-Environment In Vitro Co-Culture Model
by
Yilin Zhang, Rachel C. Anderson, Chunping You, Ajitpal Purba, Minghui Yan, Paul Maclean, Zhenmin Liu and Dulantha Ulluwishewa
Microorganisms 2024, 12(5), 873; https://doi.org/10.3390/microorganisms12050873 - 26 Apr 2024
Abstract
Intestinal barrier hyperpermeability, which is characterised by impaired tight junction proteins, is associated with a variety of gastrointestinal and systemic diseases. Therefore, maintaining intestinal barrier integrity is considered one of the effective strategies to reduce the risk of such disorders. This study aims
[...] Read more.
Intestinal barrier hyperpermeability, which is characterised by impaired tight junction proteins, is associated with a variety of gastrointestinal and systemic diseases. Therefore, maintaining intestinal barrier integrity is considered one of the effective strategies to reduce the risk of such disorders. This study aims to investigate the potential benefits of two probiotic strains (Lactiplantibacillus plantarum ST-III and Lacticaseibacillus rhamnosus KF7) on intestinal barrier function by using a physiologically relevant in vitro model of the intestinal epithelium. Our results demonstrate that both strains increased transepithelial electrical resistance, a measure of intestinal barrier integrity. Immunolocalisation studies indicated that this improvement in barrier function was not due to changes in the co-localisation of the tight junction (TJ) proteins ZO-1 and occludin. However, we observed several modifications in TJ-related genes in response to the probiotics, including the upregulation of transmembrane and cytosolic TJ proteins, as well as TJ signalling proteins. Gene expression modulation was strain- and time-dependent, with a greater number of differentially expressed genes and higher fold-change being observed in the L. plantarum ST-III group and at the latter timepoint. Further studies to investigate how the observed gene expression changes can lead to enhanced barrier function will aid in the development of probiotic foods to help improve intestinal barrier function.
Full article
(This article belongs to the Special Issue The Impact of Probiotics on Gut Health)
Open AccessArticle
Bacteria–Fungi Interactions in Multiple Sclerosis
by
Miriam Gorostidi-Aicua, Iraia Reparaz, Ane Otaegui-Chivite, Koldo García, Leire Romarate, Amaya Álvarez de Arcaya, Idoia Mendiburu, Maialen Arruti, Tamara Castillo-Triviño, Laura Moles and David Otaegui
Microorganisms 2024, 12(5), 872; https://doi.org/10.3390/microorganisms12050872 - 26 Apr 2024
Abstract
Multiple sclerosis (MS) arises from a complex interplay between host genetic factors and environmental components, with the gut microbiota emerging as a key area of investigation. In the current study, we used ion torrent sequencing to delve into the bacteriome (bacterial microbiota) and
[...] Read more.
Multiple sclerosis (MS) arises from a complex interplay between host genetic factors and environmental components, with the gut microbiota emerging as a key area of investigation. In the current study, we used ion torrent sequencing to delve into the bacteriome (bacterial microbiota) and mycobiome (fungal microbiota) of people with MS (pwMS), and compared them to healthy controls (HC). Through principal coordinate, diversity, and abundance analyses, as well as clustering and cross-kingdom microbial correlation assessments, we uncovered significant differences in the microbial profiles between pwMS and HC. Elevated levels of the fungus Torulaspora and the bacterial family Enterobacteriaceae were observed in pwMS, whereas beneficial bacterial taxa, such as Prevotelladaceae and Dialister, were reduced. Notably, clustering analysis revealed overlapping patterns in the bacteriome and mycobiome data for 74% of the participants, with weakened cross-kingdom interactions evident in the altered microbiota of pwMS. Our findings highlight the dysbiosis of both bacterial and fungal microbiota in MS, characterized by shifts in biodiversity and composition. Furthermore, the distinct disease-associated pattern of fungi–bacteria interactions suggests that fungi, in addition to bacteria, contribute to the pathogenesis of MS. Overall, our study sheds light on the intricate microbial dynamics underlying MS, paving the way for further investigation into the potential therapeutic targeting of the gut microbiota in MS management.
Full article
(This article belongs to the Section Gut Microbiota)
►▼
Show Figures
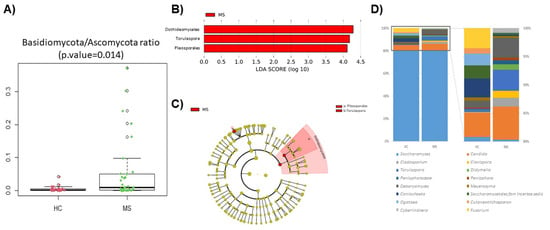
Figure 1
Open AccessArticle
Gender Impacted Gut Microbiota and Growth Performance in the Blotched Snakehead (Channa maculata)
by
Chang Fang, Fang Zeng, Shijun Chen, Shuisheng Li, Yuting Yang, Wanjing Lin, Yun Liu, Cheng Peng and Huirong Yang
Microorganisms 2024, 12(5), 871; https://doi.org/10.3390/microorganisms12050871 - 26 Apr 2024
Abstract
The blotched snakehead Channa maculata is an important economical freshwater species in East Asia. However, there has been relatively little research conducted on the correlation between gender and gut microbes. In this study, 36 of 1000 blotched snakeheads were randomly selected for growth
[...] Read more.
The blotched snakehead Channa maculata is an important economical freshwater species in East Asia. However, there has been relatively little research conducted on the correlation between gender and gut microbes. In this study, 36 of 1000 blotched snakeheads were randomly selected for growth performance measurement and gut microbiota high-throughput sequencing. Results showed that microbial diversity, composition, and metabolic functions were altered by gender and growth performance except the microbial network. In our study, Proteobacteria were the most abundant phylum, with Fusobacteria showing enrichment in males and Bacteroidetes in females. Notably, phylum Deinococcus-Thermus was identified as a significant biomarker. The Cetobacterium was the most abundant genus-level taxon. Furthermore, gut microbes specializing in the production of gut-healthy substances, such as coenzymes and vitamins, were identified as biomarkers in the fast-growing group. Our investigation highlighted the impact of gender on the composition and abundance of gut microbial biomarkers in both males and females, thereby influencing differential growth performance through the modulation of specific metabolic functions.
Full article
(This article belongs to the Special Issue Beneficial Microorganisms in Aquaculture)
►▼
Show Figures
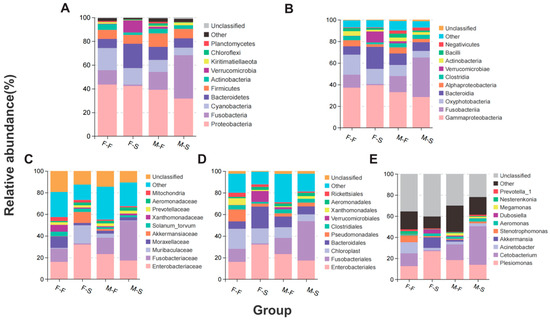
Figure 1
Open AccessArticle
Plant Growth-Promoting Bacteria Influence Microbial Community Composition and Metabolic Function to Enhance the Efficiency of Hybrid pennisetum Remediation in Cadmium-Contaminated Soil
by
Zhao-Jin Chen, Meng-Lu Li, Shan-Shan Gao, Yu-Bo Sun, Hui Han, Bai-Lian Li and Yu-Ying Li
Microorganisms 2024, 12(5), 870; https://doi.org/10.3390/microorganisms12050870 - 26 Apr 2024
Abstract
The green and efficient remediation of soil cadmium (Cd) is an urgent task, and plant-microbial joint remediation has become a research hotspot due to its advantages. High-throughput sequencing and metabolomics have technical advantages in analyzing the microbiological mechanism of plant growth-promoting bacteria in
[...] Read more.
The green and efficient remediation of soil cadmium (Cd) is an urgent task, and plant-microbial joint remediation has become a research hotspot due to its advantages. High-throughput sequencing and metabolomics have technical advantages in analyzing the microbiological mechanism of plant growth-promoting bacteria in improving phytoremediation of soil heavy metal pollution. In this experiment, a pot trial was conducted to investigate the effects of inoculating the plant growth-promoting bacterium Enterobacter sp. VY on the growth and Cd remediation efficiency of the energy plant Hybrid pennisetum. The test strain VY-1 was analyzed using high-throughput sequencing and metabolomics to assess its effects on microbial community composition and metabolic function. The results demonstrated that Enterobacter sp. VY-1 effectively mitigated Cd stress on Hybrid pennisetum, resulting in increased plant biomass, Cd accumulation, and translocation factor, thereby enhancing phytoremediation efficiency. Analysis of soil physical-chemical properties revealed that strain VY-1 could increase soil total nitrogen, total phosphorus, available phosphorus, and available potassium content. Principal coordinate analysis (PCoA) indicated that strain VY-1 significantly influenced bacterial community composition, with Proteobacteria, Firmicutes, Chloroflexi, among others, being the main differential taxa. Redundancy analysis (RDA) revealed that available phosphorus, available potassium, and pH were the primary factors affecting bacterial communities. Partial Least Squares Discriminant Analysis (PLS-DA) demonstrated that strain VY-1 modulated the metabolite profile of Hybrid pennisetum rhizosphere soil, with 27 differential metabolites showing significant differences, including 19 up-regulated and eight down-regulated expressions. These differentially expressed metabolites were primarily involved in metabolism and environmental information processing, encompassing pathways such as glutamine and glutamate metabolism, α-linolenic acid metabolism, pyrimidine metabolism, and purine metabolism. This study utilized 16S rRNA high-throughput sequencing and metabolomics technology to investigate the impact of the plant growth-promoting bacterium Enterobacter sp. VY-1 on the growth and Cd enrichment of Hybrid pennisetum, providing insights into the regulatory role of plant growth-promoting bacteria in microbial community structure and metabolic function, thereby improving the microbiological mechanisms of phytoremediation.
Full article
(This article belongs to the Topic Microbe-Induced Abiotic Stress Alleviation in Plants)
►▼
Show Figures

Figure 1
Open AccessArticle
The Contributions of Sub-Communities to the Assembly Process and Ecological Mechanisms of Bacterial Communities along the Cotton Soil–Root Continuum Niche Gradient
by
Shaodong Liu, Ruihua Liu, Siping Zhang, Qian Shen, Jing Chen, Huijuan Ma, Changwei Ge, Lidong Hao, Jinshan Zhang, Shubing Shi and Chaoyou Pang
Microorganisms 2024, 12(5), 869; https://doi.org/10.3390/microorganisms12050869 (registering DOI) - 26 Apr 2024
Abstract
Soil microbes are crucial in shaping the root-associated microbial communities. In this study, we analyzed the effect of the soil–root niche gradient on the diversity, composition, and assembly of the bacterial community and co-occurrence network of two cotton varieties. The results revealed that
[...] Read more.
Soil microbes are crucial in shaping the root-associated microbial communities. In this study, we analyzed the effect of the soil–root niche gradient on the diversity, composition, and assembly of the bacterial community and co-occurrence network of two cotton varieties. The results revealed that the bacterial communities in cotton soil–root compartment niches exhibited a skewed species abundance distribution, dominated by abundant taxa showing a strong spatial specificity. The assembly processes of the rhizosphere bacterial communities were mainly driven by stochastic processes, dominated by the enrichment pattern and supplemented by the depletion pattern to recruit bacteria from the bulk soil, resulting in a more stable bacterial community. The assembly processes of the endosphere bacterial communities were determined by processes dominated by the depletion pattern and supplemented by the enrichment pattern to recruit species from the rhizosphere, resulting in a decrease in the stability and complexity of the community co-occurrence network. The compartment niche shaped the diversity of the bacterial communities, and the cotton variety genotype was an important source of diversity in bacterial communities within the compartment niche. We suggest that the moderate taxa contribute to significantly more changes in the diversity of the bacterial community than the rare and abundant taxa during the succession of bacterial communities in the cotton root–soil continuum.
Full article
(This article belongs to the Section Plant Microbe Interactions)
►▼
Show Figures
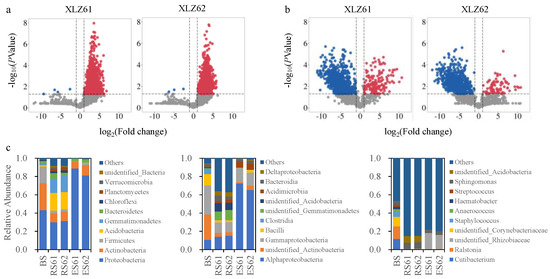
Figure 1

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, Ecologies, Forests, Microorganisms, Plants
Litter Decompositions: From Individuals to Ecosystems
Topic Editors: Wen Zhou, Guihua LiuDeadline: 30 April 2024
Topic in
Aquaculture Journal, Fishes, Microorganisms, Water
Women in Aquaculture Research
Topic Editors: Camino Ordás, Patrícia Díaz-RosalesDeadline: 30 June 2024
Topic in
Agronomy, Environments, Microorganisms, Pollutants, Sustainability, Water
Soil and Water Pollution Process and Remediation Technologies, 2nd Volume
Topic Editors: Hongbiao Cui, Ru Wang, Yu Shi, Haiying Lu, Lin ChenDeadline: 15 July 2024
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024

Conferences
Special Issues
Special Issue in
Microorganisms
Plant and Human Probiotics: Consequences on the Autochthonous Microbiota
Guest Editors: Francesco Vuolo, Elisa Gamalero, Elisa Bona, Giorgia NovelloDeadline: 30 April 2024
Special Issue in
Microorganisms
Human Skin Microbiota 2.0
Guest Editors: Holger Brüggemann, Rolf LoodDeadline: 15 May 2024
Special Issue in
Microorganisms
The Urban Microbiome
Guest Editors: Haruo Suzuki, Soojin JangDeadline: 31 May 2024
Special Issue in
Microorganisms
Emerging Pathogens Causing Acute Hepatitis
Guest Editors: Ibrahim M. Sayed, Sayed F. Abdelwahab, Ahmed El-ShamyDeadline: 15 June 2024
Topical Collections
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde
Topical Collection in
Microorganisms
Biodegradation and Environmental Microbiomes
Collection Editors: Shuangjiang Liu, Hongzhi Tang, Jiandong Jiang, Xiaolei Wu